Personnel de l'université
Dorian MCILROY
Maître de ConférenceCoordonnées
CR2TI UMR1064 CHU Nantes - Hôtel Dieu 30 Bd Jean Monnet 44093 Nantes Cedex 01 Téléphone 02 44 76 81 82
- Dorian.Mcilroy@univ-nantes.fr
- Site internet
- 30 Bd Jean Monnet
Discipline(s) enseignée(s)
Virologie (L3, M1), Microbiologie Générale (L2), Mécanismes d'Evolution (L1)
Thèmes de recherche
Neutralizing antibody response to BK polyomavirus (BKPyV); evolution of the BKPyV capsid.
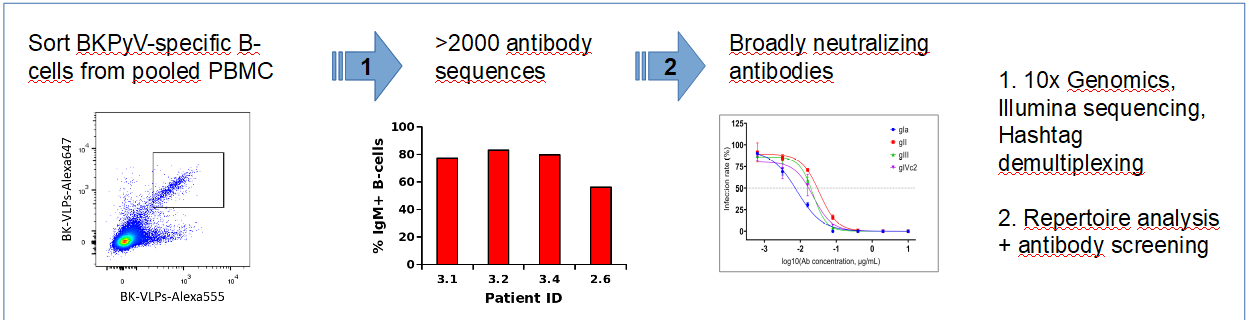
With respect to antibodies, in the context of the ANR BKNAB project, Ngoc-Khanh Nguyen (PhD co-directed with X. Saulquin CRCINA, defended 31/3/2021) was able to sort BKPyV-specific B-cells using labeled BKPyV capsids, and analyze their antibody repertoire by 10xGenomics single-cell RNASeq. This allowed us to generate several neutralizing monoclonal antibodies against the virus.
Data from this project was presented in oral communications at national (AFIPP 2019, SFI 2021) and international (IUMS 2020) scientific meetings.
Publication - Nguyen et al. 2023
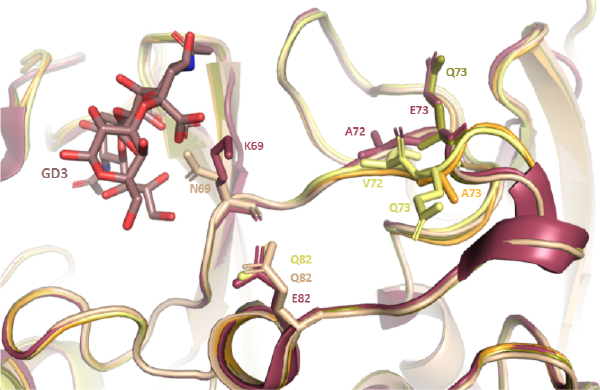
In kidney transplant recipients with persistent BKPyV replication, we found that mutations in the capsid can allow the virus to escape from the host antibody response, and also modify the tropism of the virus. Recently, Marie Sorin (PhD co-directed with Thilo Stehle, Tubingen University, defended 15/19/2022) obtained the crystal structures of several of these mutants, including one variant that does not bind to any known BKPyV receptor, but is nevertheless infectious.
Data presented in oral communications at AFIPP 2021.
Publication Sorin et al. 2023
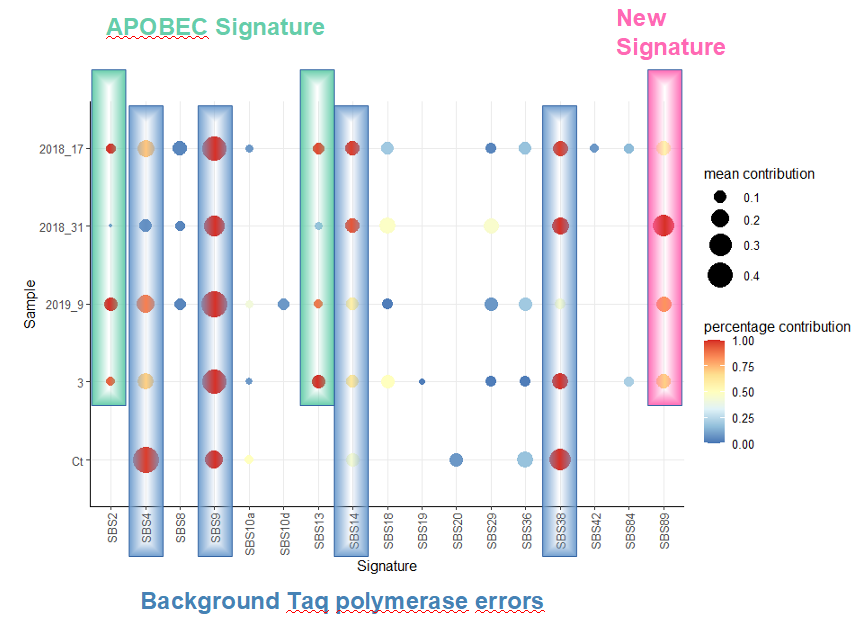
To understand how mutations arise in the virus genome, Nicolas Le Baut (Master 1 project, 2021) analyzed NGS data from patient samples to identify patterns in the types of mutations that arise in the VP1 gene. This confirmed that editing by the host APOBEC proteins is a major driver of virus genome mutation, and we also identified a separate mutation signature, suggesting that at least one other biological process contributes to the genetic variability of BKPyV.
Publication - McIlroy et al. 2022

With respect to antibodies, in the context of the ANR BKNAB project, Ngoc-Khanh Nguyen (PhD co-directed with X. Saulquin CRCINA, defended 31/3/2021) was able to sort BKPyV-specific B-cells using labeled BKPyV capsids, and analyze their antibody repertoire by 10xGenomics single-cell RNASeq. This allowed us to generate several neutralizing monoclonal antibodies against the virus.
Data from this project was presented in oral communications at national (AFIPP 2019, SFI 2021) and international (IUMS 2020) scientific meetings.
Publication - Nguyen et al. 2023

In kidney transplant recipients with persistent BKPyV replication, we found that mutations in the capsid can allow the virus to escape from the host antibody response, and also modify the tropism of the virus. Recently, Marie Sorin (PhD co-directed with Thilo Stehle, Tubingen University, defended 15/19/2022) obtained the crystal structures of several of these mutants, including one variant that does not bind to any known BKPyV receptor, but is nevertheless infectious.
Data presented in oral communications at AFIPP 2021.
Publication Sorin et al. 2023

To understand how mutations arise in the virus genome, Nicolas Le Baut (Master 1 project, 2021) analyzed NGS data from patient samples to identify patterns in the types of mutations that arise in the VP1 gene. This confirmed that editing by the host APOBEC proteins is a major driver of virus genome mutation, and we also identified a separate mutation signature, suggesting that at least one other biological process contributes to the genetic variability of BKPyV.
Publication - McIlroy et al. 2022
Informations complémentaires
Recent publications:
Structural and functional analysis of natural capsid variants suggests sialic acid-independent entry of BK polyomavirus.
Sorin MN, Di Maio A, Silva LM, Ebert D, Delannoy CP, Nguyen NK, Guerardel Y, Chai W, Halary F, Renaudin-Autain K, Liu Y, Bressollette-Bodin C, Stehle T, McIlroy D.Cell Rep. 2023 Feb 14;42(2):112114. doi: 10.1016/j.celrep.2023.112114. PMID: 36790933
A cluster of broadly neutralizing IgG against BK polyomavirus in a repertoire dominated by IgM.
Nguyen NK, Devilder MC, Gautreau-Rolland L, Fourgeux C, Sinha D, Poschmann J, Hourmant M, Bressollette-Bodin C, Saulquin X, McIlroy D.Life Sci Alliance. 2023 Jan 30;6(4):e202201567. doi: 10.26508/lsa.202201567. PMID: 36717250
Quantification of APOBEC3 Mutation Rates Affecting the VP1 Gene of BK Polyomavirus In Vivo.
McIlroy D, Peltier C, Nguyen ML, Manceau L, Mobuchon L, Le Baut N, Nguyen NK, Tran MC, Nguyen TC, Bressollette-Bodin C.
Viruses. 2022 Sep 19;14(9):2077. doi: 10.3390/v14092077.
PMID: 36146883
Persistent BK Polyomavirus Viruria is Associated with Accumulation of VP1 Mutations and Neutralization Escape.
McIlroy D, Hönemann M, Nguyen NK, Barbier P, Peltier C, Rodallec A, Halary F, Przyrowski E, Liebert U, Hourmant M, Bressollette-Bodin C.
Viruses. 2020 Jul 29;12(8):824. doi: 10.3390/v12080824.PMID: 32751274
Intra-patient viral evolution in polyomavirus-related diseases.
McIlroy D, Halary F, Bressollette-Bodin C.Philos Trans R Soc Lond B Biol Sci. 2019 May 27;374(1773):20180301. doi: 10.1098/rstb.2018.0301.PMID: 30955497
Structural and functional analysis of natural capsid variants suggests sialic acid-independent entry of BK polyomavirus.
Sorin MN, Di Maio A, Silva LM, Ebert D, Delannoy CP, Nguyen NK, Guerardel Y, Chai W, Halary F, Renaudin-Autain K, Liu Y, Bressollette-Bodin C, Stehle T, McIlroy D.Cell Rep. 2023 Feb 14;42(2):112114. doi: 10.1016/j.celrep.2023.112114. PMID: 36790933
A cluster of broadly neutralizing IgG against BK polyomavirus in a repertoire dominated by IgM.
Nguyen NK, Devilder MC, Gautreau-Rolland L, Fourgeux C, Sinha D, Poschmann J, Hourmant M, Bressollette-Bodin C, Saulquin X, McIlroy D.Life Sci Alliance. 2023 Jan 30;6(4):e202201567. doi: 10.26508/lsa.202201567. PMID: 36717250
Quantification of APOBEC3 Mutation Rates Affecting the VP1 Gene of BK Polyomavirus In Vivo.
McIlroy D, Peltier C, Nguyen ML, Manceau L, Mobuchon L, Le Baut N, Nguyen NK, Tran MC, Nguyen TC, Bressollette-Bodin C.
Viruses. 2022 Sep 19;14(9):2077. doi: 10.3390/v14092077.
PMID: 36146883
Persistent BK Polyomavirus Viruria is Associated with Accumulation of VP1 Mutations and Neutralization Escape.
McIlroy D, Hönemann M, Nguyen NK, Barbier P, Peltier C, Rodallec A, Halary F, Przyrowski E, Liebert U, Hourmant M, Bressollette-Bodin C.
Viruses. 2020 Jul 29;12(8):824. doi: 10.3390/v12080824.PMID: 32751274
Intra-patient viral evolution in polyomavirus-related diseases.
McIlroy D, Halary F, Bressollette-Bodin C.Philos Trans R Soc Lond B Biol Sci. 2019 May 27;374(1773):20180301. doi: 10.1098/rstb.2018.0301.PMID: 30955497
Mis à jour le 17 avril 2026.

